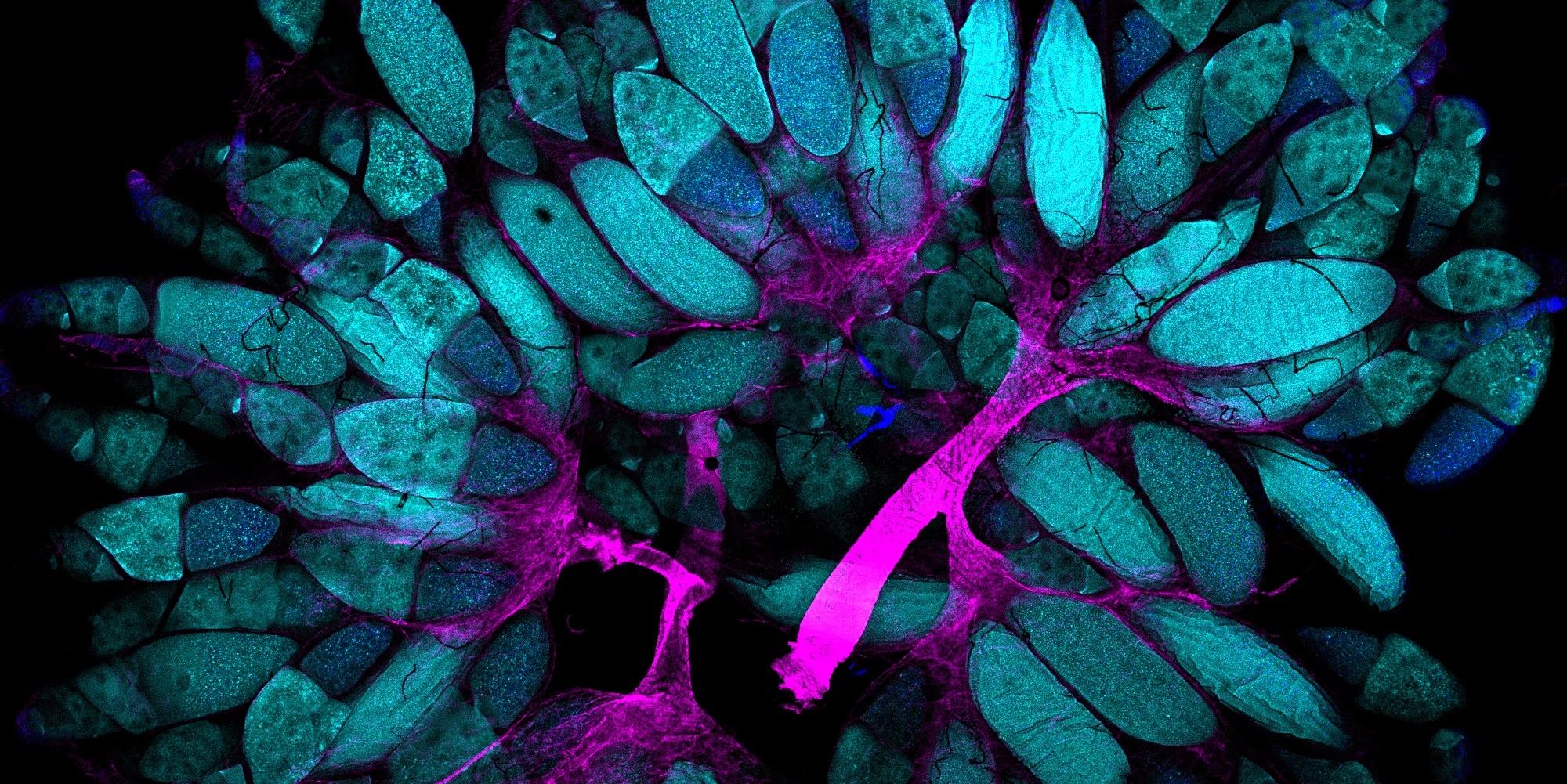
For more information please visit: WeilLabCambridge.com
Regulating protein expression is fundamental to the life of any cell. One way this can be done is through mRNA association with biomolecular condensates, membrane-less compartments that function as reaction crucibles and sub-cellular organisational hubs. Condensates have been shown to be involved in effectively all aspects of biology, including development, neurobiology, and pathogenesis, and are a promising target for new therapeutic innovation. However, to inform future drug discovery, an in vivo molecular understanding of condensates is required.
In early animal development, condensates are important in the patterning of embryonic axes, formation of neuronal networks, and movement of cells. We test the formation and function of a conserved biomolecular condensate, processing bodies (P Bodies), in Drosophila exploiting their advantages for sophisticated genetics and live imaging. Research in fruit flies has identified thousands of genes with human homologues and has provided key insights into developmental pathways, oncology, neurobiology and immunology.
Our use of the egg chamber and early embryo has the experimental benefits of being an in vivo developing system, allowing for a controllable switch at egg activation, ease of physical manipulation, and providing ample material for experimentation. Recently, we have shown that P bodies in the mature Drosophila egg are primarily regulated by structurally distinct proteins and weak multivalent interactions. In vivo, P body integrity is controlled through an arrested physical state which is critical for the storage of mRNAs.
Overall, we aim to fully elucidate the formation, maintenance and dynamics of P bodies and how these condensates control mRNA metabolism. This work will shed light on the fundamental mechanisms controlling cellular protein production and inform medical advancements.
